
icon16 is here! Learn how we’re expanding AutoNorm™ access to all sequencing labs in this press release
Learn moreApplication Note
Brochure
Case Study
Technical Note
Application Note
Application Note
Technical Note
Application Note
Application Note
Brochure
Application Note

Why Library Quality Is Set During Amplification — Not After. Your sequencing pool looks perfect on the Bioanalyzer. Every sample reads at ~10 nM. The gel lane is clean. You're ready to load. But here's the uncomfortable truth: equal concentration is not equal information. A balanced pool built on over-amplified, complexity-collapsed libraries isn't a sequencing success — it's an expensive illusion.
Read more
Penn State genomics technologist Kerry Hair put iconPCR with AutoNorm through its paces — and uncovered a paradigm shift in amplicon sequencing, microbiome profiling, and index behavior. Discover what's hiding at the margins of your workflow.
Read more
NGS library prep generates more plastic waste than most labs realize. Learn how icon96 and icon16 eliminate redundant workflow steps, reduce consumables by up to 95%, and help your lab go greener without compromising data quality.
Read more
Discover how n6's iconPCR with AutoNorm technology won Best New Product, expanded to 30+ countries, and partnered with genomics leaders in 2025's breakthrough year.
Read moreCompare Normalase vs Normalizer vs AutoNorm for NGS library normalization. iconPCR™ eliminates separate normalization steps and prevents over-amplification artifacts—no beads, no math, no extra reagents needed.
Read more
iconPCR™ eliminates NGS library quantification, saving labs 20-40% prep time and $7-20 per sample. No qPCR plates, no normalization steps — just perfectly balanced libraries from PCR.
Read moreWhen it comes to wrangling the wild world of RNA-seq library prep, especially for those notorious FFPE samples, n6 gets it—science is demanding, but your workflow can be less so. That’s why pairing NEB kits with the n6 iconPCR 96-well thermocycler and AutoNorm technology is a match made for sequencing success.
Read more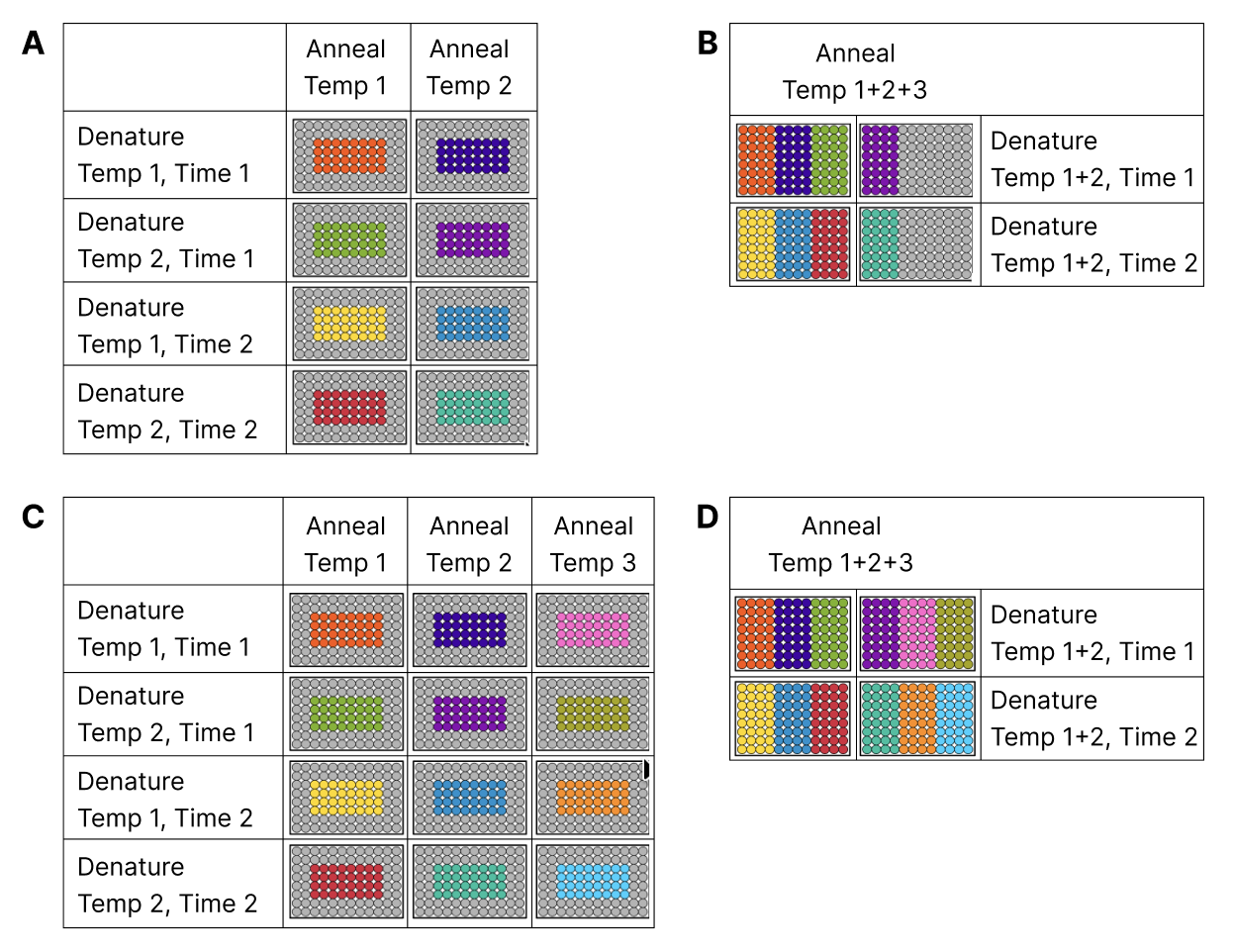
Assay development is at the core of scientific discovery in genomics, diagnostics, and life science. Yet, optimizing PCR workflows has always meant battling with split-plot constraints, endless thermocycler runs, and mountains of consumable waste. If you’re seeking a game-changer for your lab, it's time to meet iconPCR with AutoNorm™—the benchtop thermocycler reimagined by n6 for the next-generation scientist.
Read more
The landscape of metagenomics and microbiome research is evolving at breakneck speed. As sequencing technologies improve and the demand for high-resolution, bias-free microbial data grow...
Read more
In the rapidly evolving field of genomics, advancing technology continues to redefine research capabilities...
Read moreWhen a next-generation sequencing run fails, it's not just a minor laboratory setback. It's the cost of a new car—literally. With all-in costs ranging from $30,000 to $120,000 from sample to answer on platforms like Illumina's...
Read more
When it comes to NGS library prep, there’s one step that can make or break your sequencing run: NGS library...
Read more
Welcome to the wild world of genomics, where science meets serious problem-solving—and plenty of innovation. The Center forAdvanced Technology (CAT) at UCSF...
Read moreCase Study
bioRxiv
Geib, S. et al. (2026)
See detailsbioRxiv
Green, S. et al. (2026)
See detailsbioRxiv
Mason, C.J. et al. (2026)
See detailsbioRxiv
Jouvenot, Y. et al. (2024)
See detailsABRF 2026
Jouvenot Y, Austin W, Arrastia M, Patel P.
n6, Pleasanton, California, 94566
SLAS 2026
Sushant Khanal1, Yann Jouvenot2, Vanessa Process1, Alaina Villarreal1, Luca Beker1, Kelly Buckman2, Martina Werner1, & Ulrich Thomann1
1) Covaris LLC, Woburn, MA, USA 2) n6, Pleasanton, California, USA
SLAS 2026
Jouvenot Y1, Austin W1, Echave P2, Law W2, Obermoeller D2, Patterson S2
1) n6, Pleasanton, California, 94566 2) Revvity, Waltham, MA, 02451
PAG 2026
Austin W1, Szeluga N2, Jouvenot Y1, Hargarten H3, Ortiz-Uriarte B3, Haase A3, Haskell C4, Honaas L3, Harkess A2, Patel P1
1) n6, Pleasanton, California 2) HudsonAlpha Institute for Biotechnology, Huntsville, AL 3) USDA ARS, Wenatchee, WA 4) Washington State University, Pullman, WA
view posterASHG 2025
Austin W, Jouvenot Y, Patel P n6, Pleasanton, California, 94566
view posterASHG 2025
Jouvenot Y, Austin W, Patel P n6, Pleasanton, California, 94566
view posterCancer Genomics Consortium Annual Meeting 2025
Austin W, Jouvenot Y, Patel P n6, Pleasanton, California, 94566
view posterAGBT 2025
Jouvenot Y, Patel P n6, Pleasanton, California, 94566
view posterAGBT 2025
Jouvenot Y, Patel P n6, Pleasanton, California, 94566
view posterAGBT Ag 2025
Brett Hale1, Caroline Obert2, Kyle Metcalfe2, Andrew Boddicker2, Marielle Krivet2, Junhua Zhao2, Tuval Ben-Yehezkel2, Pranav Patel3, Yann Jouvenot3, Wes Austin3, Gagneet Kaur3
1) AgriGro, Inc, Doniphan, MO, USA 2) Element Biosciences, San Diego, CA3) n6, Pleasanton, CA
view posterGet answers to common questions about iconPCR™ technology, AutoNorm™, and how our platform can streamline your NGS workflow.
Our application specialists are here to help you optimize your NGS workflow with iconPCR technology.