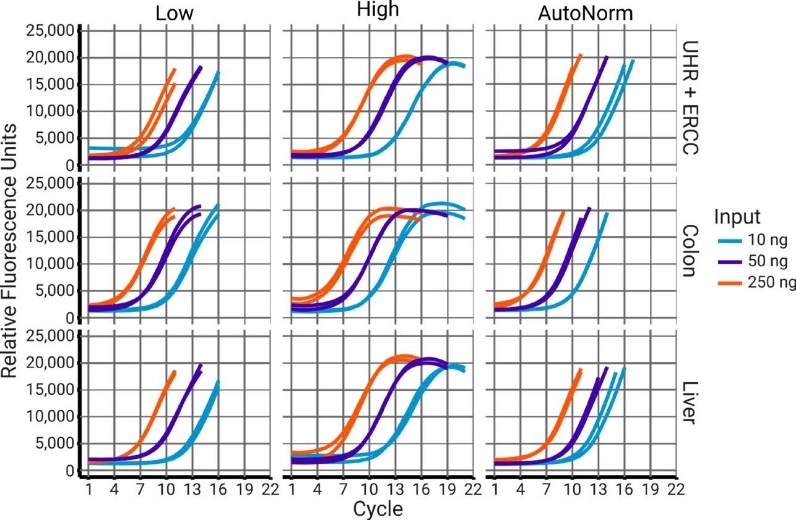
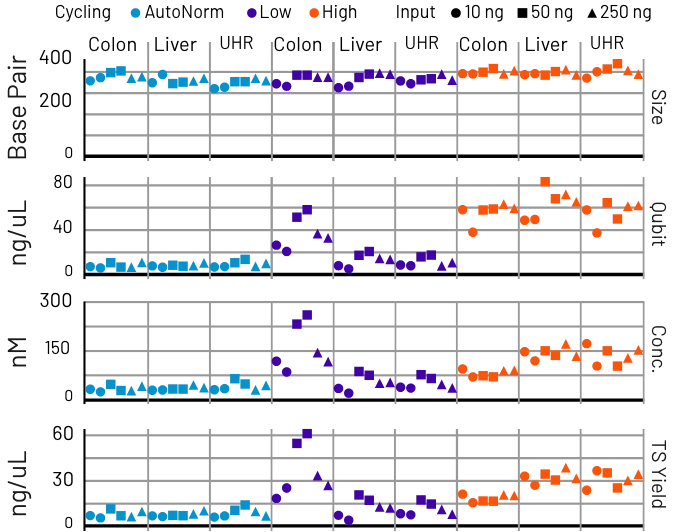
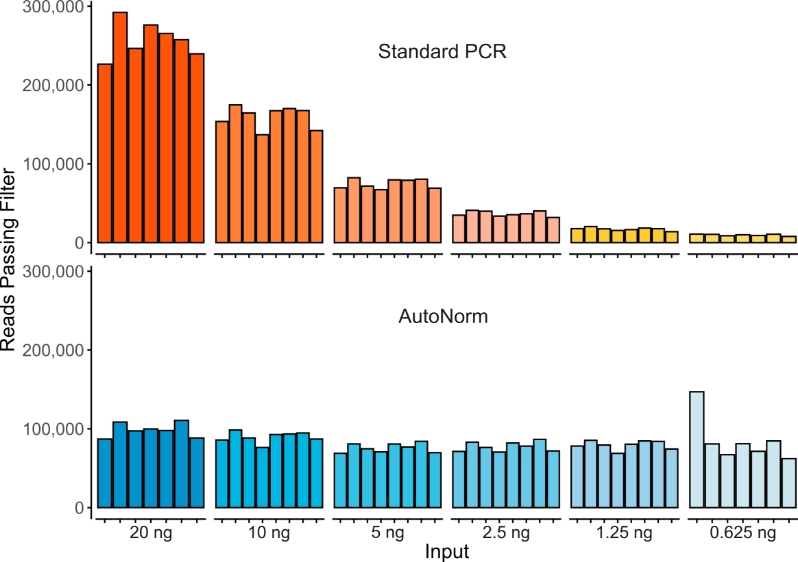
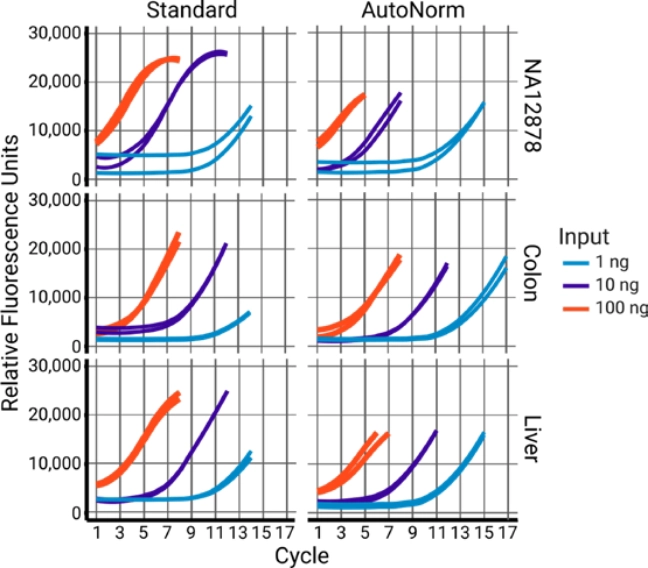
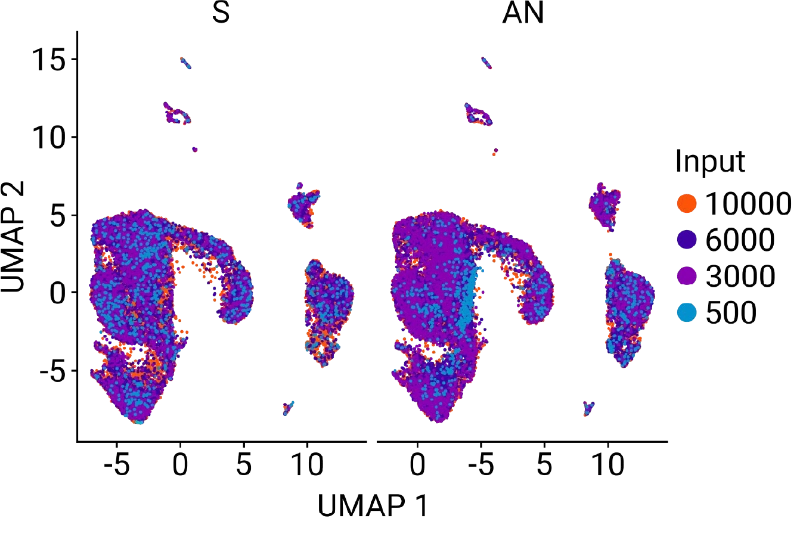
AutoNorm Flips the Script on Library Amplification
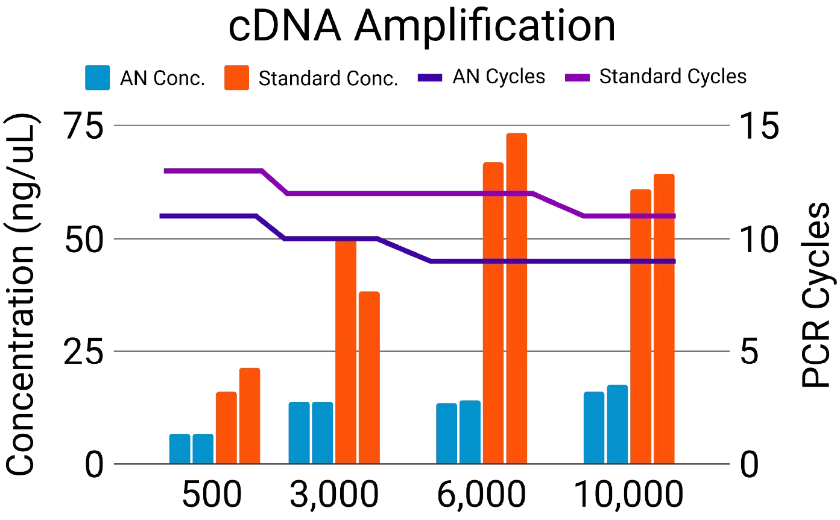
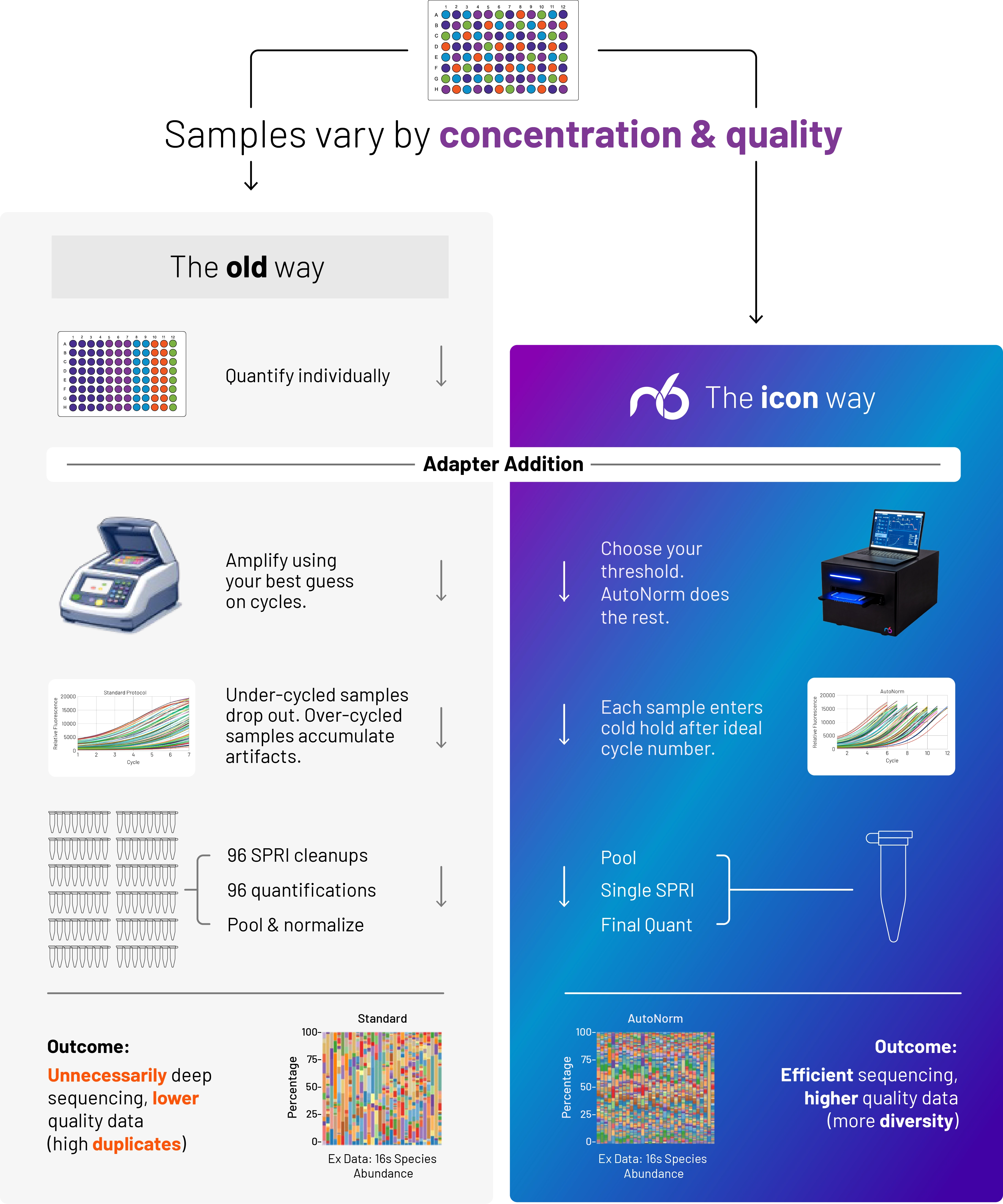
The old way: Set amplification cycles.
DNA concentration follows (sort of).
Setup
Estimate optimal amplification cycles.
Outcome
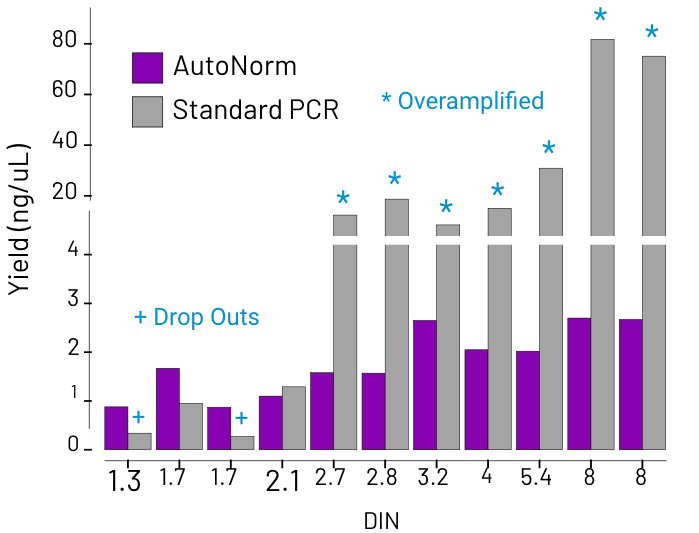
Concentration varies widely, resulting in dropout, overamplification, and the need for additional normalization.
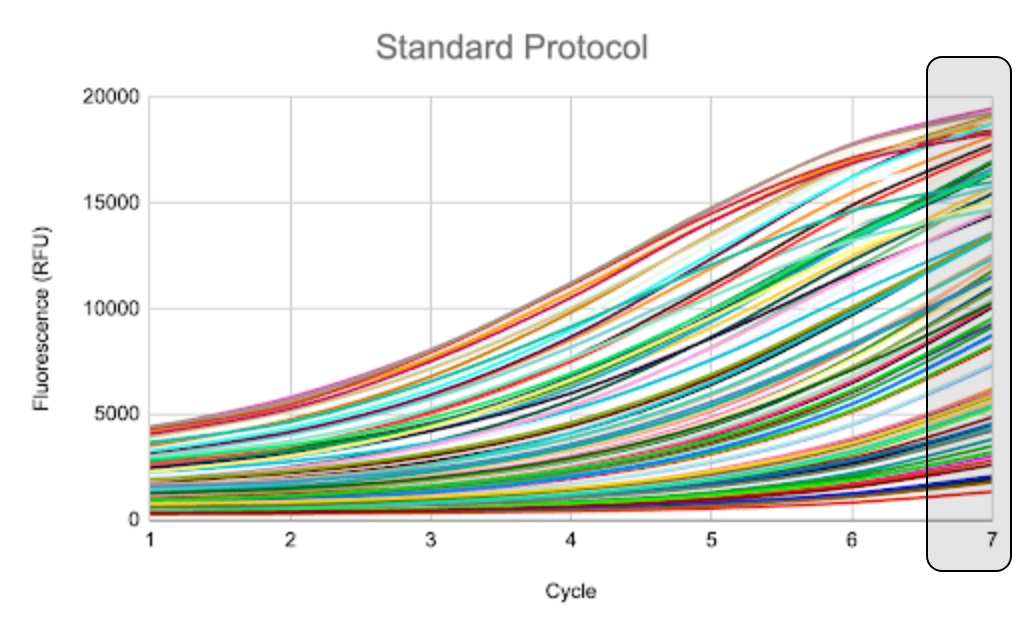
The icon way: Set target concentration.
Cycle # follows.
Setup
Set target concentration.
Outcome
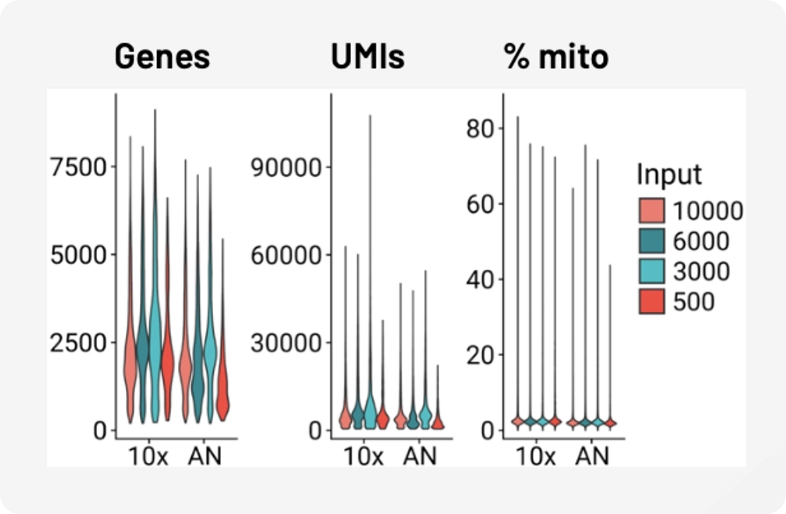
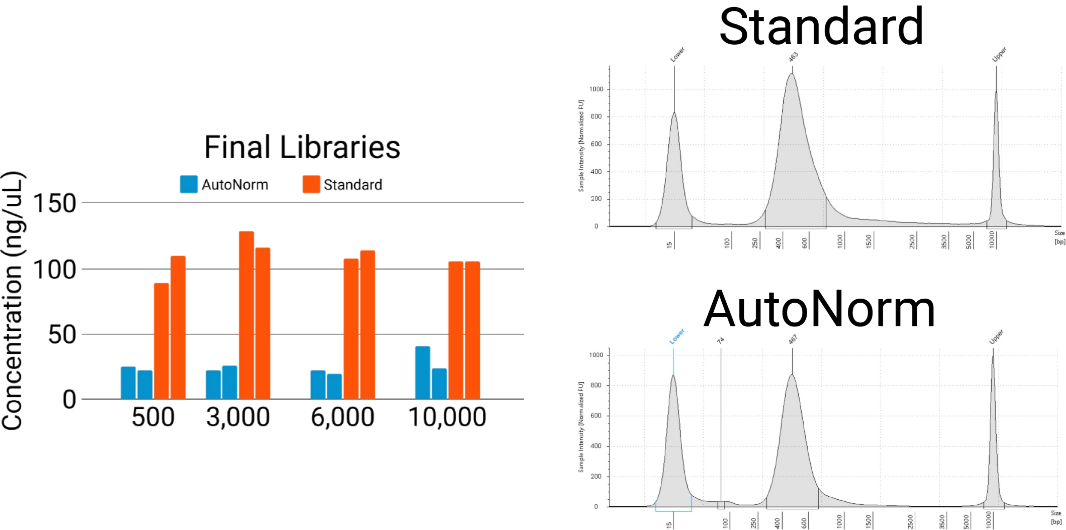
Samples are consistent and auto-optimized, no manual normalization needed.
.webp)
How it Works
First, we built a better box
Each well of the icon instrument features an individual, miniaturized thermocycler with real-time temperature monitoring.
Then, we made it smart
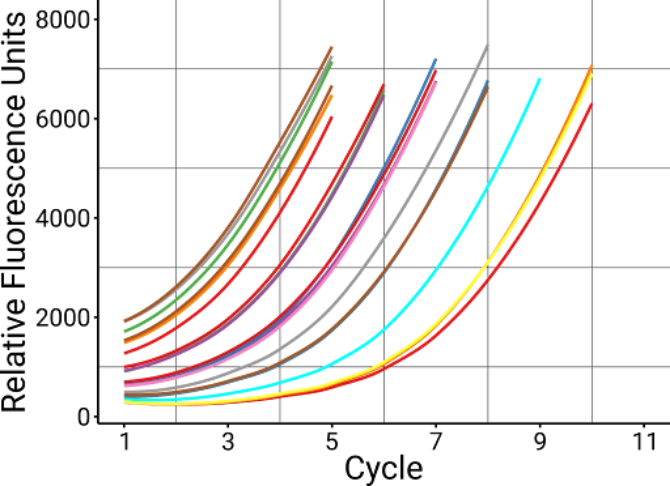
AutoNorm™ intelligent software automates cycle control by monitoring concentration in real time using fluorescence, and placing each well on a cold hold at just the right time.
Impact
Save time

Save money

Save the world

Simpler Workflow

“iconPCR is key to improving our NGS workflows — eliminating QC and normalization while preventing over amplification”
60% Reduction in hand-on time

Built-in normalization collapses three PCR runs and dozens of QC steps into a single, automated plate.
99% Faster QC

Treat 96 libraries as one: clean once (SPRI cleanup), quantify once (Qubit quant), then sequence.
Lower Costs

“By incorporating iconPCR, we can remove several QC steps that are pretty expensive… Having the opportunity to do the normalization removes that cost as well.”
0 Manual normalization

Running samples as they are and pooling immediately after amplification dramatically reduces labor costs.
50% Reduction in cost

AutoNorm-enabled sample pooling will cut library prep spend for amplicon sequencing in half while improving data quality.
Better Data

“We really do think this is an instrument that every Genomics Core should want to have… PCR is your enemy, and this is your best weapon against running too many PCR cycles, and I think we’ll just see improved data quality coming out of reducing the number of PCR cycles before you go into sequencing.”
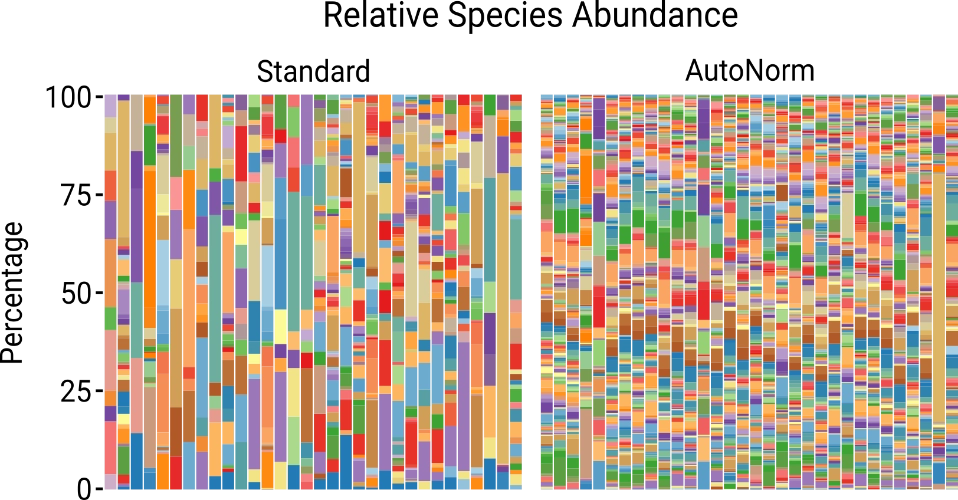
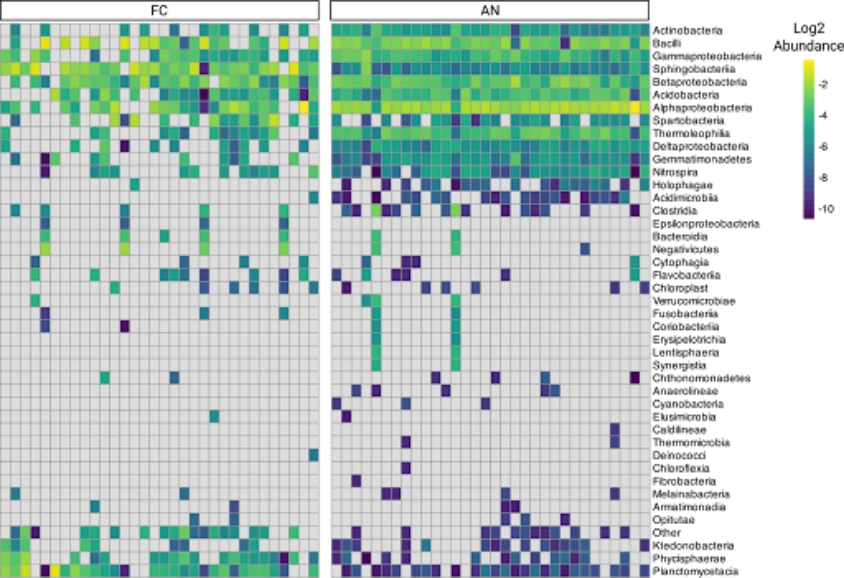
4X Fewer chimeras

In library prep for 16s metagenomic sequencing, iconPCR produced 400% less chimeras, resulting in more usable reads and a truer view of complex microbial communities.
9 Fewer PCR cycles

In full-length 16s sequencing, AutoNormalization with iconPCR entered a cold hold up to 9 cycles earlier (average), resulting in fewer artifacts.